Scientist Spotlight: Meet David Wentworth

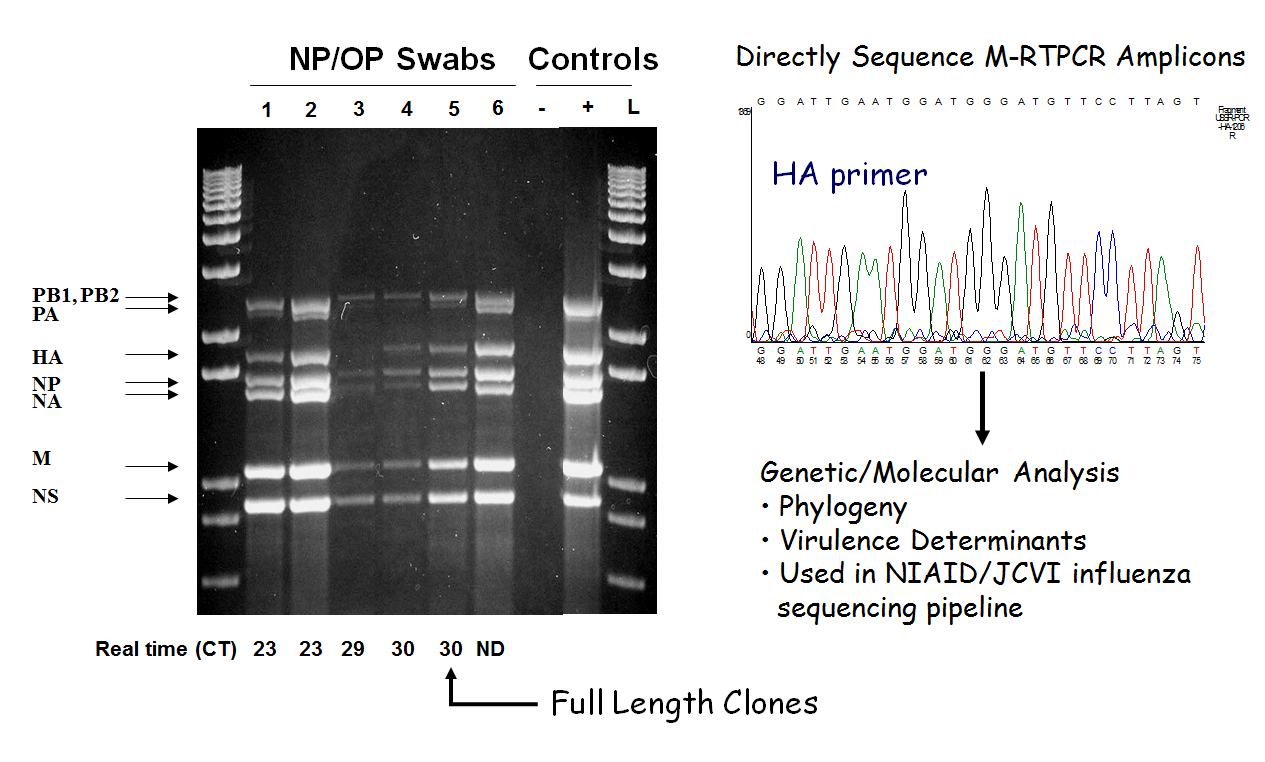
During the height of the H1N1 Flu pandemic, David Wentworth was running a microbial genetics laboratory at the Wadsworth Center, New York State Department of Health (NYSDOH) where he was instrumental in developing a method to amplify influenza genomes regardless of strain using “universal primers” or short strands of DNA that recognize conserved segments across the genomes of many different flu strains. This amplification process was developed to generate recombinant influenza A viruses (the most common flu type affecting humans and animals) that could be used for the production of new vaccines. From a clinical swab it took his team 9-12 days to develop vaccine seed stocks. It was this work that first brought Dave to JCVI’s attention.
Several years ago Dave began collaborations with JCVI scientists to sequence human and avian influenza viruses. The collaborations intensified two years ago when all pandemic flu samples (or suspected flu samples) were first sent to Dave’s lab so the virus could be amplified in sufficient quantities for sequencing using his new amplification pipeline. The amplification took only a day and then isolated, non-infectious, DNA was sent to JCVI for sequencing. JCVI was the natural choice for this work since we are host to the government-funded “Influenza Genome Sequencing Project,” with the goal of sequencing large numbers of viral genomes to help scientists worldwide to understand how flu viruses evolve and cause disease. JCVI researchers then deposited influenza sequences into GenBank within two days of receiving DNA from Dave’s lab, enabling researchers worldwide to track what strains are circulating and how they are evolving. JCVI has sequenced over 75% of the influenza genomes in GenBank, the NIH public repository for sharing genetic sequencing data.
Dave was soon invited for a talk at JCVI. “The opportunities at JCVI were to help build the [viral genomics] program. And already good, quality people are here studying viruses with a focus on viral evolution and sequencing analysis,” Dave remarked. “Being part of generating that information, I think makes you have a better feel for the biology.” The capabilities for viral sequencing combined with IFX strengths and the interest in viral evolution at JCVI was a draw for Dave and he soon joined the team. Moreover, there are opportunities at JCVI to work with collaborators who send specimens from various regions of the world for sequencing so that we can “more deeply understand the mutations that contribute to virulence,” he said. He is particularly interested in antigenic drift (how viruses escape immunity) that contributes to the “annual influenza escape,” which is critical in developing vaccine strains.
The need for new and improved methods to develop vaccines, coupled with the advances in synthetic genomics developed at JCVI led to the formation last year by JCVI and the company Synthetic Genomics Inc. of a new company, Synthetic Genomics Vaccines Inc. (SGVI). JCVI scientists, through SGVI, are working on a three-year collaboration agreement with Novartis to apply synthetic genomics tools and technologies to accelerate the production of the influenza seed strains required for vaccine manufacturing. The agreement, supported by an award from the U.S. Biomedical Advanced Research and Development Authority (BARDA), could ultimately lead to a more timely and effective response to seasonal and pandemic influenza outbreaks. The idea is to create viruses de novo or synthesize genes critical for its antigenicity and put these in normal vaccine strains for production. The goal of the work at SGVI is to synthesize a virus in one week, or rather a seed stock, which still needs to be amplified in big fermenters. New seed stocks take 3-4 weeks to produce which is currently a rate liming step.
You don’t hear too many people singing its praises and saying “I love the flu!” as Dave has remarked, but put in context, his enthusiasm for his work shines through best when talking about his love of teaching. He gets excited teaching young scientists about virology, especially helping them to understand the important areas to study, and where the research will lead to solve a major problem. “The rewarding part of being a mentor is to see all of the people who have found their niche – it might not be bench research but they are still carrying knowledge with them.”
Aside from spending time with his family, Dave enjoys a hobby started by his dad – to cultivate honey bees. A community gardens group at a middle school in Albany, NY was looking for bees to pollinate their plants. Dave spearheaded the effort and used it as a learning tool for kids, who helped feed honey to caterpillars and moths. He also used to give lectures on bee cultivation and has taught college courses in animal science. Dave’s enthusiasm for science among his students and peers could be considered infectious, just like the subject of his research!